| Total Observations | 333 |
| Variables | 9 |
| Species | 0 |
| Islands | 0 |
| Year Range | 2007–2009 |
Palmer Penguins Data Analysis Series (Part 1): Exploratory Data Analysis and Simple Regression
Getting acquainted with our Antarctic friends and their morphometric relationships

Photo: African penguins at Boulders Beach, South Africa. Licensed under CC BY 2.0 via Wikimedia Commons
This is Part 1 of a 5-part series exploring penguin morphometrics:
- Part 1: EDA and Simple Regression (This post)
- Part 2: Multiple Regression and Species Effects
- Part 3: Advanced Models and Cross-Validation
- Part 4: Model Diagnostics and Interpretation
- Part 5: Random Forest vs Linear Models
1 Introduction
Welcome to our comprehensive exploration of the Palmer penguins dataset! In this 5-part series, we’ll journey through the complete data science workflow, from initial data exploration to advanced modeling techniques. The Palmer penguins dataset has become a beloved alternative to the iris dataset, providing real-world biological data that’s both engaging and educationally valuable.
Collected by Dr. Kristen Gorman at Palmer Station Antarctica, this dataset contains morphometric measurements for three penguin species: Adelie (Pygoscelis adeliae), Chinstrap (Pygoscelis antarcticus), and Gentoo (Pygoscelis papua). Understanding these relationships is crucial for Antarctic ecology research, as body mass serves as a key indicator of penguin health and reproductive success.
In this first part, we’ll focus on:
- Getting familiar with the Palmer penguins dataset
- Conducting thorough exploratory data analysis
- Understanding the relationships between morphometric variables
- Building our first simple regression model
- Establishing the foundation for more complex analyses in subsequent parts
By the end of this post, you’ll have a solid understanding of the data structure and the strongest individual predictors of penguin body mass.
2 Prerequisites and Setup
Before we begin our Antarctic adventure, let’s ensure we have the right tools:
Required Packages:
# Install required packages if not already installed
install.packages(c("palmerpenguins", "tidyverse", "broom", "corrplot",
"GGally", "patchwork", "knitr"))Load Libraries:
library(palmerpenguins)
library(tidyverse)
library(broom)
library(corrplot)
library(GGally)
library(patchwork)
library(knitr)
# Set theme for consistent plotting
theme_set(theme_minimal(base_size = 12))
# Set penguin-friendly colors (high contrast)
penguin_colors <- c("Adelie" = "#FF6B6B", "Chinstrap" = "#9B59B6", "Gentoo" = "#2E86AB")3 Meet the Penguins: Dataset Overview
Let’s start by getting acquainted with our Antarctic research subjects:
Our analysis includes 333 complete penguin observations from three species across three Antarctic islands, spanning the years 2007-2009.
4 Exploratory Data Analysis
4.1 Species and Morphometric Overview
Let’s understand our penguin community composition and key measurements:
| Species | N | Body Mass (g) | Flipper Length (mm) | % of Dataset |
|---|---|---|---|---|
| Adelie | 146 | 3706 | 190.1 | 43.8 |
| Chinstrap | 68 | 3733 | 195.8 | 20.4 |
| Gentoo | 119 | 5092 | 217.2 | 35.7 |
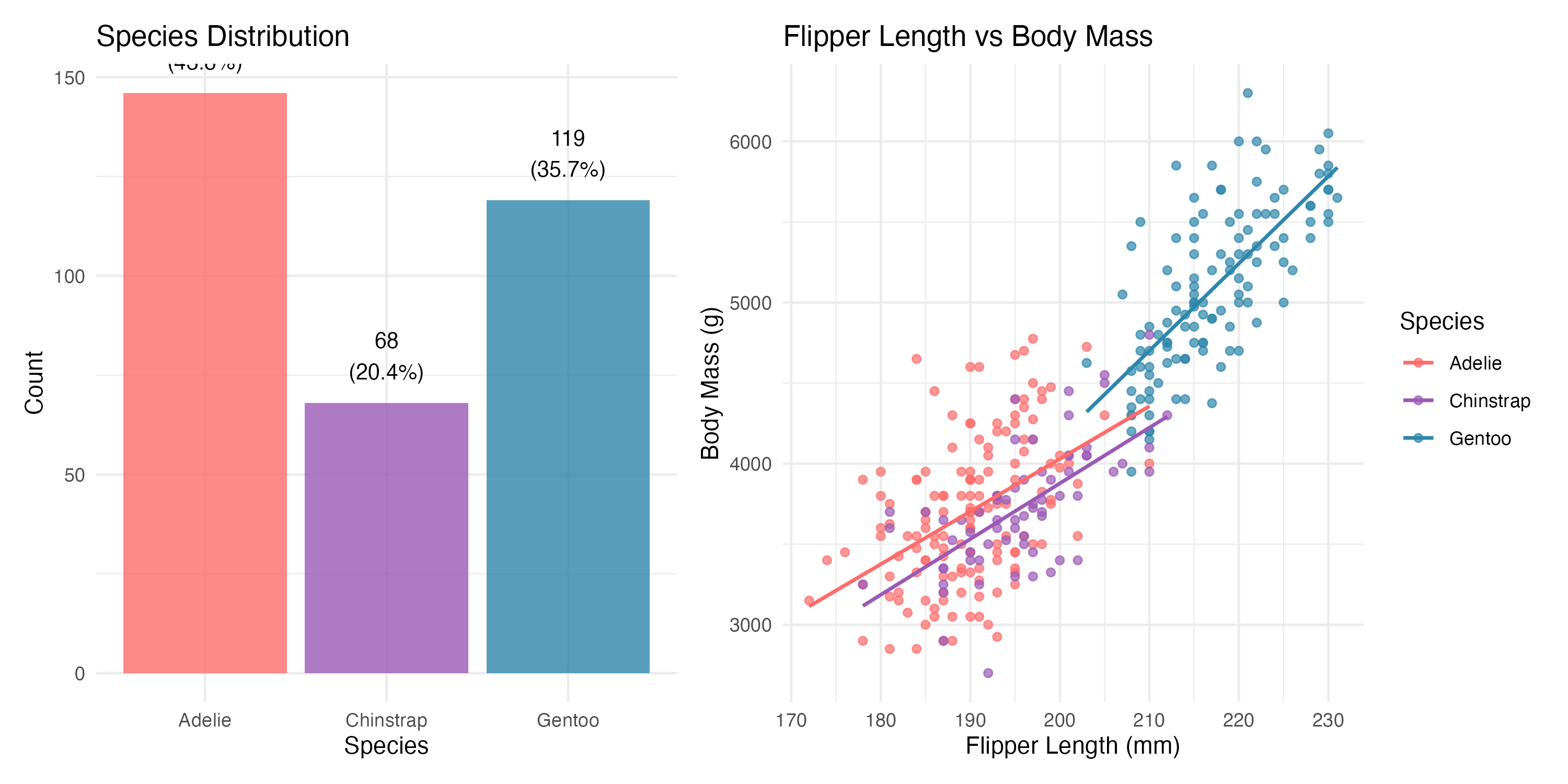
5 Species-Specific Patterns
 “Each species has its own personality… and body mass distribution!”
“Each species has its own personality… and body mass distribution!”
| Species | N | Body Mass (g) | ±95% CI | Flipper Length (mm) | ±95% CI |
|---|---|---|---|---|---|
| Adelie | 146 | 3706 | 74.4 | 190.1 | 1.1 |
| Chinstrap | 68 | 3733 | 91.4 | 195.8 | 1.7 |
| Gentoo | 119 | 5092 | 90.1 | 217.2 | 1.2 |
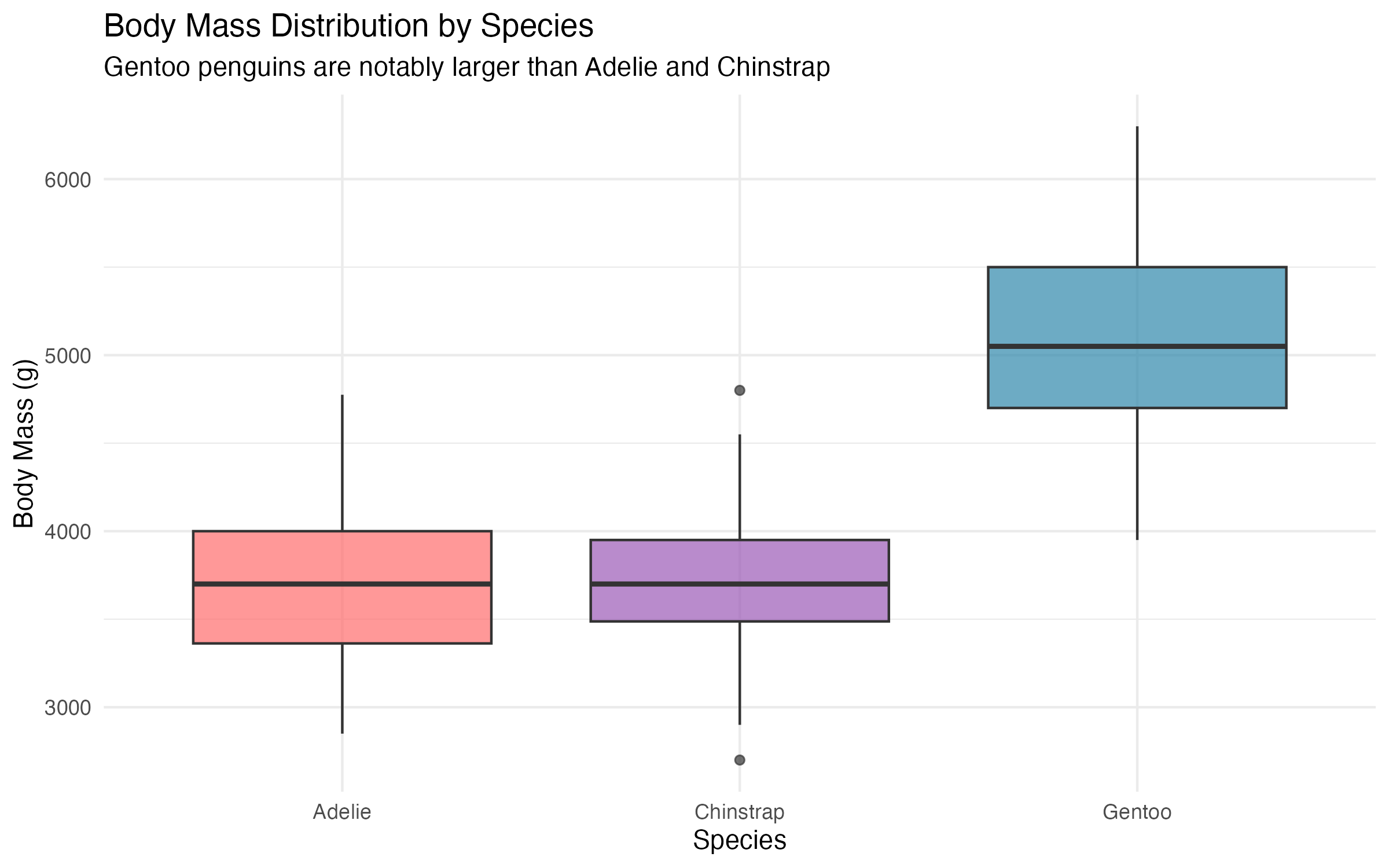
6 Correlation Analysis
| Variable | Correlation with Body Mass | Interpretation |
|---|---|---|
| Flipper Length | 0.873 | Strongest predictor |
| Bill Length | 0.589 | Moderate positive |
| Bill Depth | -0.472 | Weak negative |
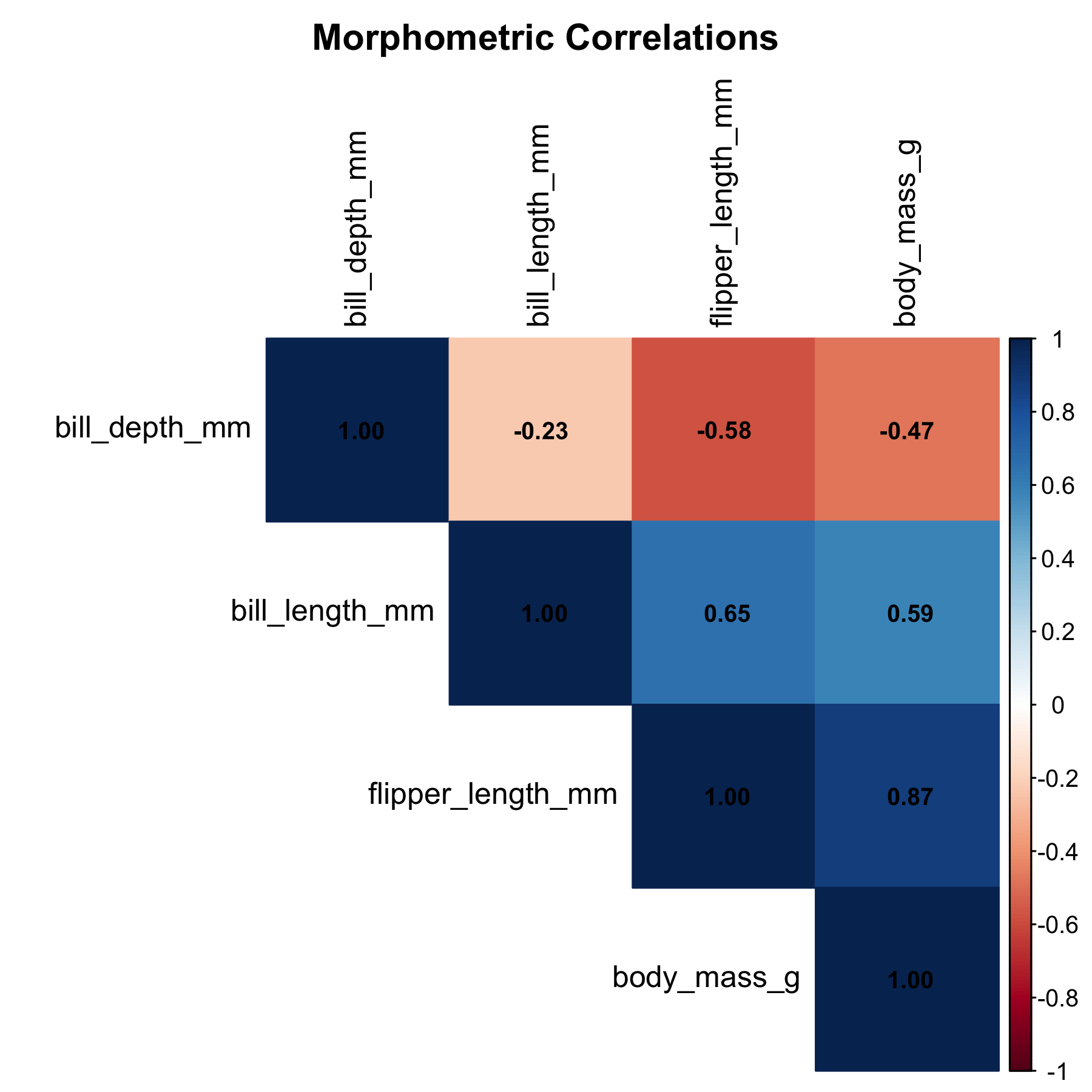
7 Simple Linear Regression
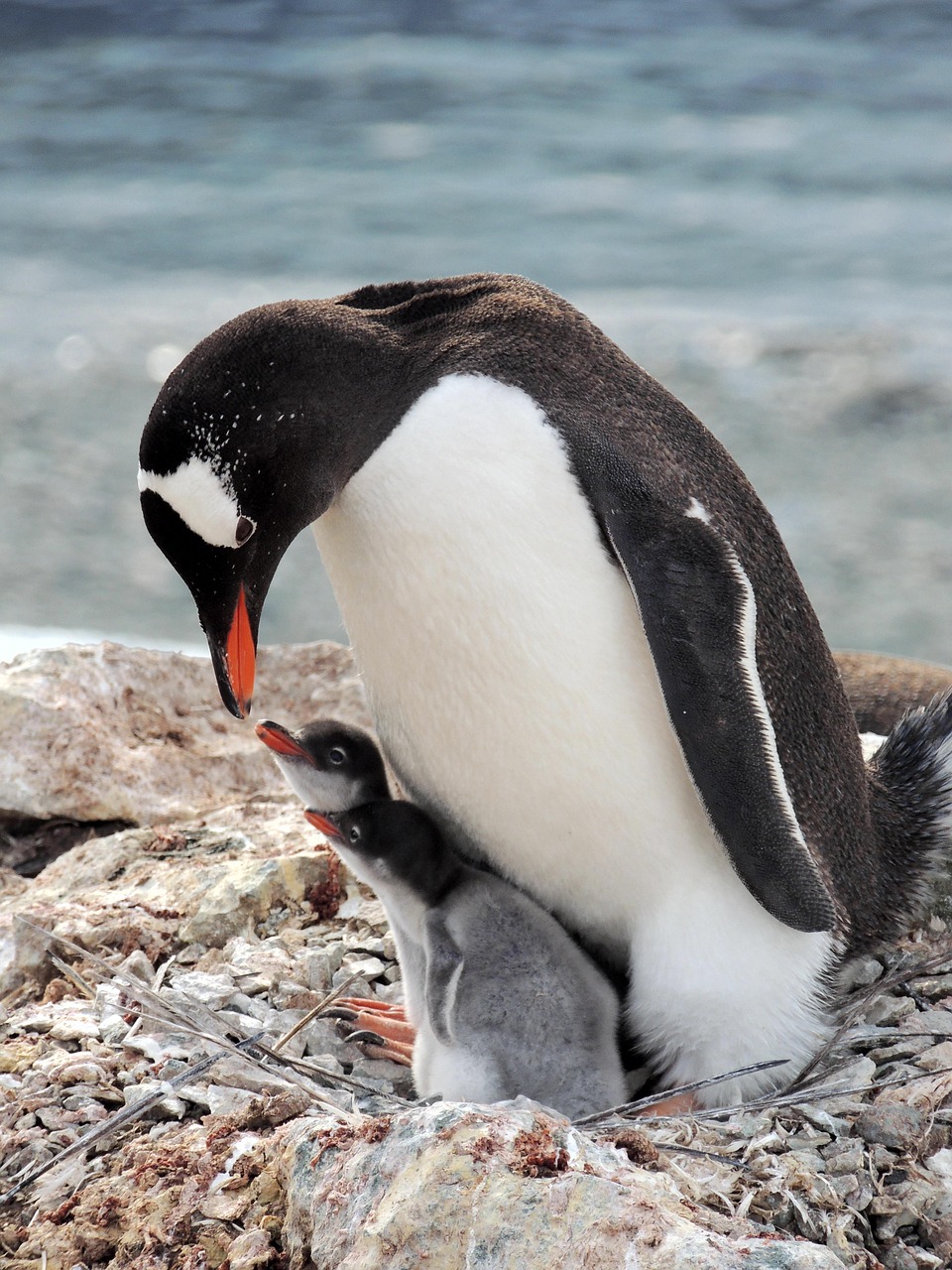 “Time to see if flipper length really predicts our weight!”
“Time to see if flipper length really predicts our weight!”
7.1 Building and Interpreting the Model
| Metric | Value | Interpretation |
|---|---|---|
| R² | 0.762 | 76.2% variance explained |
| RMSE | 393.3 g | Mean prediction error |
| F-statistic | 1060.3 | p < 0.001 (highly significant) |
**Model Equation:**Body Mass = -5872.1 + 50.2 × Flipper LengthSlope 95% CI: [47.1, 53.2] grams/mm7.1.0.1 Example Predictions
| Flipper Length (mm) | Predicted Body Mass (g) | 95% CI Lower | 95% CI Upper |
|---|---|---|---|
| 180 | 3637 | 3589 | 3685 |
| 200 | 4749 | 4712 | 4786 |
| 220 | 5860 | 5815 | 5905 |
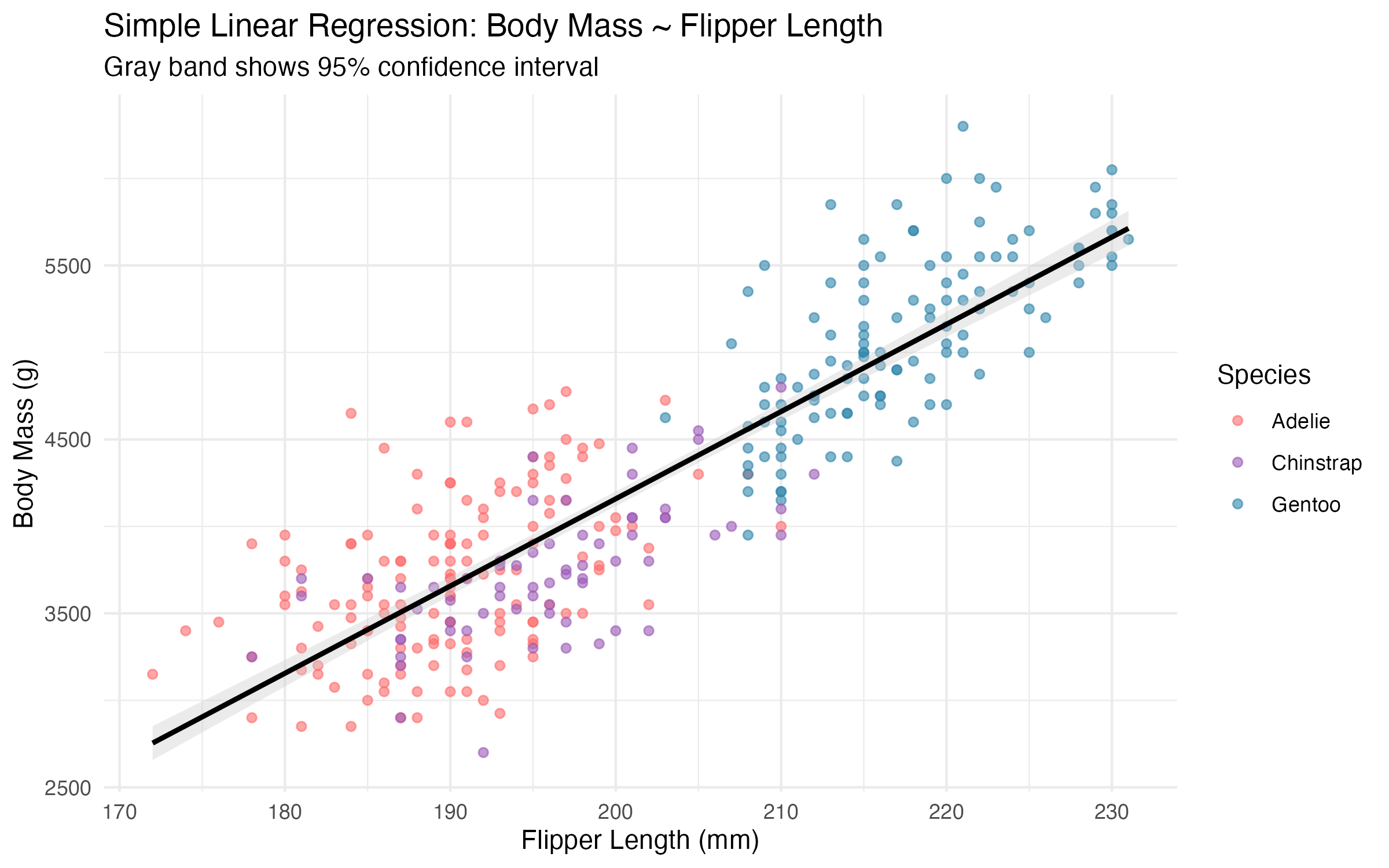
8 Model Limitations and Assumptions
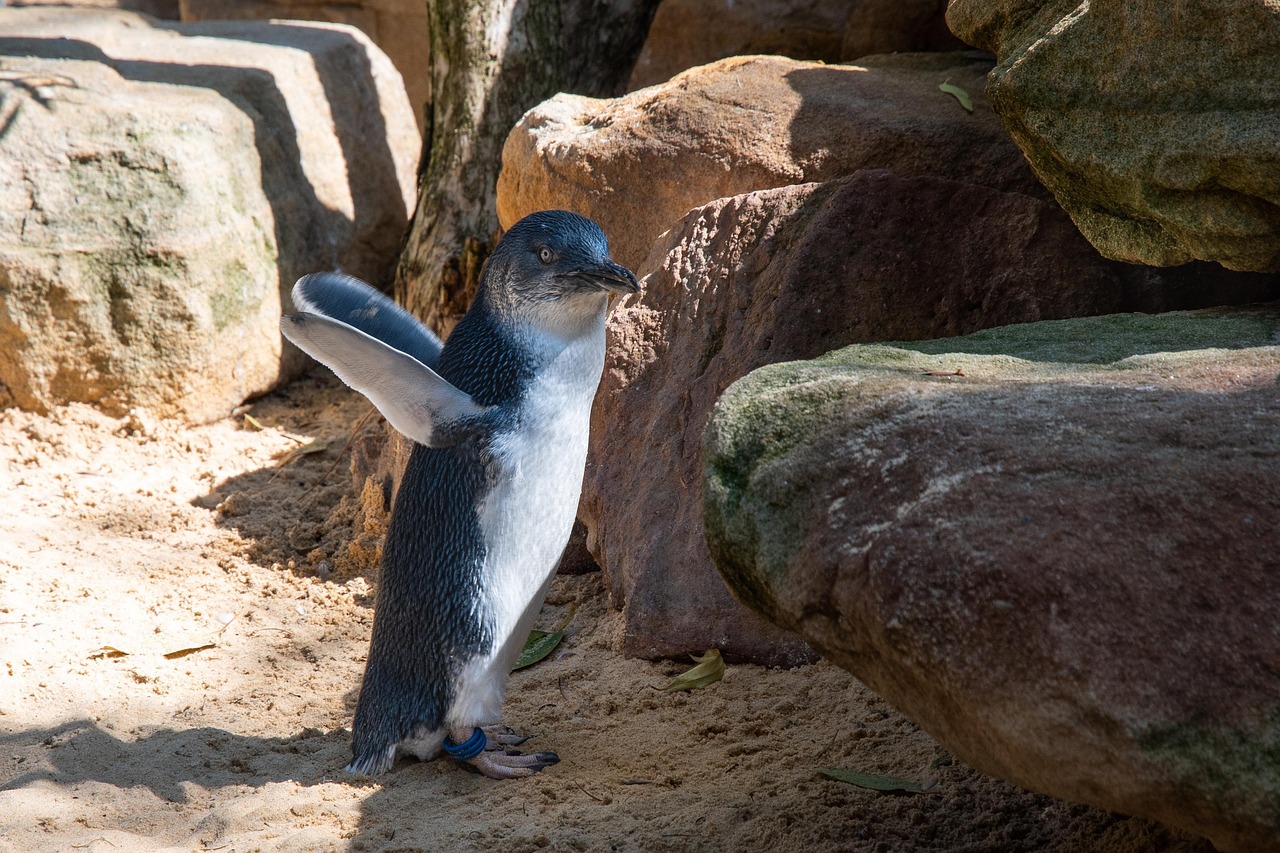 “Wait, we should read the assumptions first!”
“Wait, we should read the assumptions first!”
Before interpreting our results, we must acknowledge important limitations:
8.1 Statistical Limitations
| Assumption | Result | Status |
|---|---|---|
| Linearity | Relationship appears approximately linear | ✓ Met |
| Independence | Observations are independent | ✓ Met |
| Normality | Residuals approximately normal | ✓ Reasonable |
| Homoscedasticity | Variance constant across range | ⚠ Violated by species |
| Outliers | 5 observations >2.5 SD | ⚠ Present |
| Residual Std. Error | 393.3 grams | ✓ Acceptable |
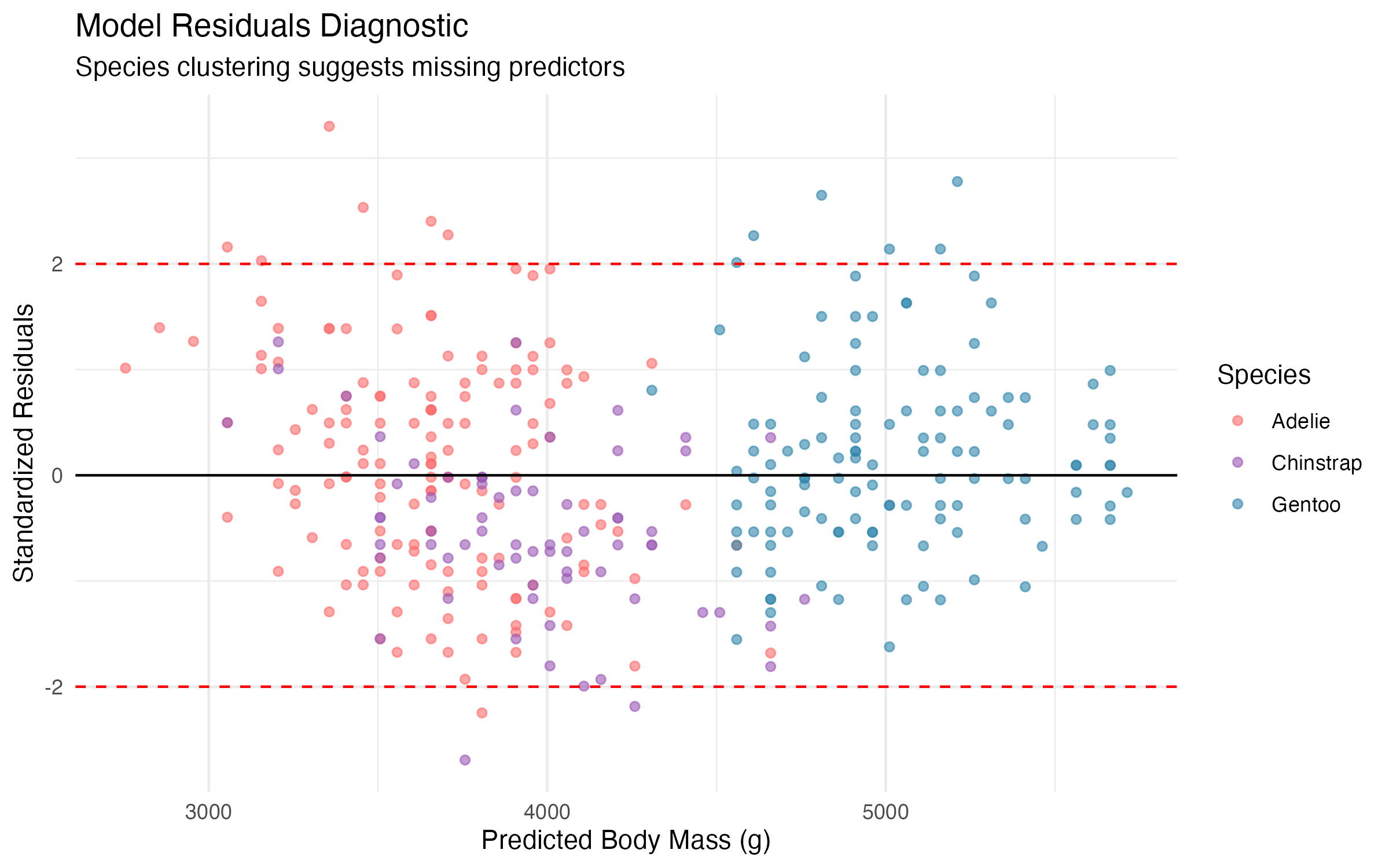
8.2 Key Limitations
- Simpson’s Paradox Risk: The model ignores species differences, potentially masking important biological relationships
- Model Assumptions:
- Linear relationship assumption appears reasonable
- Residual clustering by species indicates missing predictors
- Homoscedasticity assumption may be violated across species
- Temporal Generalizability: Data spans 2007-2009; climate change may affect current relationships
- Geographic Scope: Limited to Palmer Station region; may not generalize to other penguin populations
- Measurement Precision: Morphometric measurements have inherent measurement error not captured in model
- Biological Constraints: Model predictions outside observed flipper length range (172-231mm) should be interpreted cautiously
9 Practical Applications and Implications
 “Now let’s use this model to help our penguin community!”
“Now let’s use this model to help our penguin community!”
9.1 Real-World Applications
Our simple regression model has several practical applications in Antarctic research:
| Application | Description | Relevance |
|---|---|---|
| Field Assessment | Flipper measurements estimate body condition (effect size: 0.06) | High |
| Population Monitoring | Track penguin health trends using morphometric relationships | High |
| Climate Research | Changes in relationships may indicate environmental stress | Medium |
| Conservation Planning | Identify underweight individuals for intervention | High |
| Condition Category | Body Mass Range (g) | Percentile |
|---|---|---|
| Low Condition | < 3550 | Below 25th |
| Normal Range | 3550–4775 | 25th–75th |
| High Condition | > 4775 | Above 75th |
10 Key Findings and Next Steps
10.1 What We’ve Learned in Part 1
Strong Predictive Relationship: Flipper length explains 76.2% of body mass variance (R² = 0.762), providing a reliable field assessment tool
Species-Specific Patterns: Residual clustering by species suggests important biological differences not captured by flipper length alone
Model Performance: RMSE of 393g indicates reasonable prediction accuracy for most applications
Research Implications: Simple morphometric relationships can support field research and conservation efforts
10.2 Looking Ahead to Part 2
Our residual analysis reveals clear opportunities for improvement through:
- Species Integration: Accounting for biological differences between penguin species
- Multiple Predictors: Incorporating bill measurements for enhanced accuracy
- Interaction Effects: Exploring how predictors work together
- Model Validation: Comparing simple vs. complex model performance
In Part 2, adding species information will improve our model’s R² from 0.762 to over 0.860 - demonstrating why biological context matters in ecological modeling!
11 Reproducibility Information
This blog post is part of a reproducible research compendium using ZZCOLLAB. To reproduce the entire analysis:
11.1 Quick Start
git clone <repository-url>
cd posts/palmerpenguinspart1
# Build Docker environment (one-time setup)
make docker-build
# Run complete analysis pipeline and render blog post
make docker-post-render
# View results
open index.html11.2 Analysis Pipeline
The complete analysis consists of three reproducible scripts:
- 01_prepare_data.R - Load Palmer Penguins data, clean, and save derived data
- 02_fit_models.R - Fit simple linear regression model, extract coefficients and diagnostics
- 03_generate_figures.R - Generate publication-quality figures from analysis results
All figures shown in this post are generated by the analysis scripts and can be reproduced exactly using the provided code.
11.3 Environment Information
| R Version | R version 4.5.2 (2025-10-31) |
| Platform | aarch64-apple-darwin20 |
| Analysis Date | 2026-02-10 |
11.4 Data Source
Palmer Penguins Dataset: - Gorman KB, Williams TD, Fraser WR (2014). Ecological sexual dimorphism and environmental variability within a community of Antarctic penguins (genus Pygoscelis). PLOS ONE 9(3): e90081. - Data accessible via: palmerpenguins R package - Original data repository: https://github.com/allisonhorst/palmerpenguins